Diffuse
Updates
Jun 18, 2024
Our Vision and Initial Results
Diffuse is building generative AI for protein design.
Our mission is to build AI systems that engineer new and useful proteins with unprecedented control and accuracy.
Background
Proteins are molecules that mediate nearly all of the cellular processes that underlie life. If we can reliably engineer proteins, we can develop breakthrough therapeutics, diagnostics, and vaccines faster and with far higher rates of success than we see today.
Traditional computational protein design methods employ physics-based models. These methods have had some successes, but frequently result in designs that fail in later experiments, due to inaccuracies in the underlying models. We realized that deep learning could surpass the capabilities of these human-modeled systems, similar to advancements seen in computer vision and natural language. This idea motivated our team's early work in building generative AI for protein design.
Over the past few years, there have been rapid advancements in the application of deep learning to problems in protein engineering and design, most notably in structure prediction 1 and generative design 2. Our breakthrough results in early 2022 showed for the first time that diffusion models can design realistic and accurate protein structures.
Despite impressive computational results, we are still in the early stages of developing AI for protein design. A whole host of challenges remain - including demonstrating control over protein-protein binding, engineering larger and more complex protein structures, and enabling controllable design of novel enzymatic functions, to name a few.
Where we are today
Today we introduce Diffuse StructGen-1 (DSG-1), a new foundation model for protein design. DSG-1 can perform protein design tasks such as generating 3D protein structures based on user prompts and creating proteins to bind to a desired binding site on a target protein.
Design a binder to a target protein
Application: Binder design
We used DSG-1 to design binders - proteins that attach to target proteins at a specific region of interest. Demonstrating control over binding is an important engineering advance in protein design, as binding is a necessary step in developing protein therapeutics, reagents, and diagnostics. In this case, we asked the model to design nanobodies, a therapeutically relevant class of binders, against 2 targets.
High hit rate
Remarkably, we found that the AI-designed nanobodies bound their respective targets with a ~3% hit rate, about seven orders of magnitude better than industry-standard discovery approaches3.
Put another way, if DSG-1 produces 100 designs, there is a >95% chance at least one of them will bind to the desired target. If the standard industry approach starts with 1,000,000 candidates, there is a >99.9% chance that not a single one will bind to the desired target.
Strong and specific binding
Importantly, DSG-1 can design strong binders with affinities <=10nM - a significant advance over recent results4. Experimental data shows that DSG-1 binders (1) bind the region of interest on the target as designed and (2) do not have off-target binding. In contrast, industry-standard approaches have limited to no control over the region of binding and often produce non-specific proteins that have off-target binding.
More details can be found in our technical appendix
Going forward
We view control over binding as the first of many steps needed to advance AI for protein design. We envision a future where AI enables rapid and accurate design of any protein, from therapeutics that treat disease, to enzymes that catalyze new kinds of chemical reactions, to novel gene editing molecules.
We are an interdisciplinary team of scientists and machine learning researchers and engineers dedicated to making this future a reality.
If our mission resonates with you, join us or partner with us
- the Diffuse team
1 AlphaFold2 and 3
2 First diffusion results, RFDiffusion, Chroma
3 The standard approach to discover binders involves repeatedly panning large libraries of proteins against the target of interest. This results in a handful of "hits" that bind the target, although with little to no control over where the hits bind. The hit rate of such approaches has historically been roughly 1:1,000,000,000 (see Table 1 and Table 4 here for reference).
Appendix
Designing nanobody binders - hit rate
We asked DSG-1 to design nanobody binders at specific epitopes (regions of interest) on two target antigens. We then screened 4,000 designs per target on yeast display. We saw 3% of expressing cells in the double positive (binding and expressing) condition; in other words, 3% of the proteins that successfully expressed also bound the target (Figure 1).
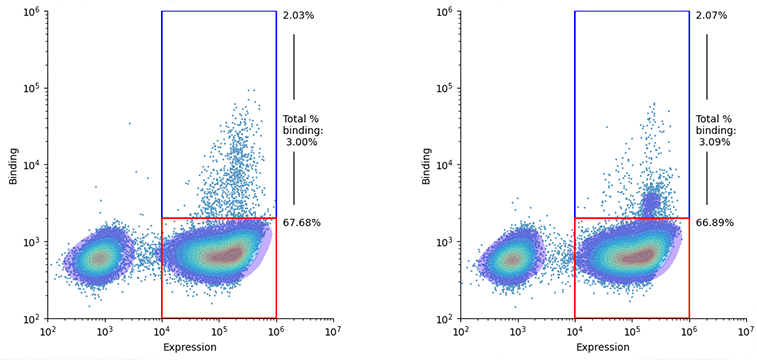
Figure 1: Screening DSG-1 designed proteins via yeast display. Points are yeast cells and kernel density estimation is overlaid. We test 4000 proteins per target. We see ~3% of the pool of proteins that successfully express on yeast (red and blue gates) also show binding to the target (blue gate). We isolate the binding population (blue), identify the proteins via next-generation sequencing, and select a subset of designs to test individually.
Binding affinity and specificity
We then characterized several proteins individually to assess their affinity to the target. Below, we plot the affinities (KDs) of all of the binders we tested (Figure 2). We see that several de novo designed nanobodies are strong binders, with affinities <= 10nM.
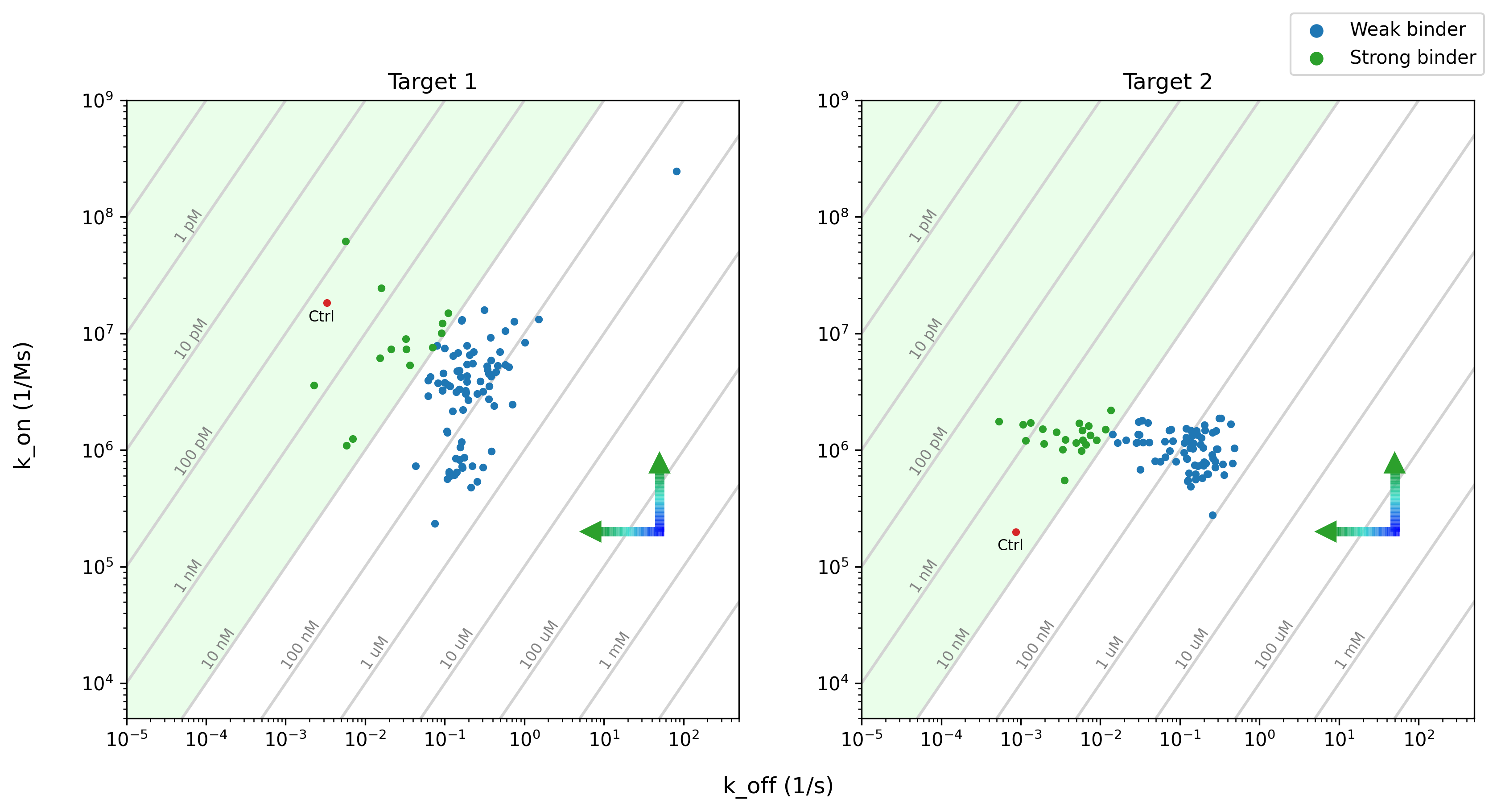
Figure 2: DSG-1 designs specific binders with a range of affinities (gray lines), including several strong binders (k_off/k_on <= 10nM, green, shaded). k_on is the association rate (how rapidly the binder binds to the target), and k_off is the dissociation rate (time for the binder to dissociate from the target). Tighter binders associate faster (up arrow) and dissociate slower (left arrow). Known binder controls shown in red.
We also confirmed that these binders are specific, and do not show off-target binding. Note that we do not use any experimental data to improve these binders - these are direct designs from the AI.
Epitope characterization
Finally, we confirmed that the DSG-1 binders bind the target epitope, as designed. For each target, we selected a known binder against the epitope of interest, as well as a second binder that binds an alternate, non-overlapping epitope on the target. If our designs bind the epitope of interest, they should be blocked from binding the target in complex with the known binder. However, they should be able to bind the target in complex with the second binder (Figure 3).
We found that every DSG-1 binder we tested binds the epitope of interest (Figure 4).
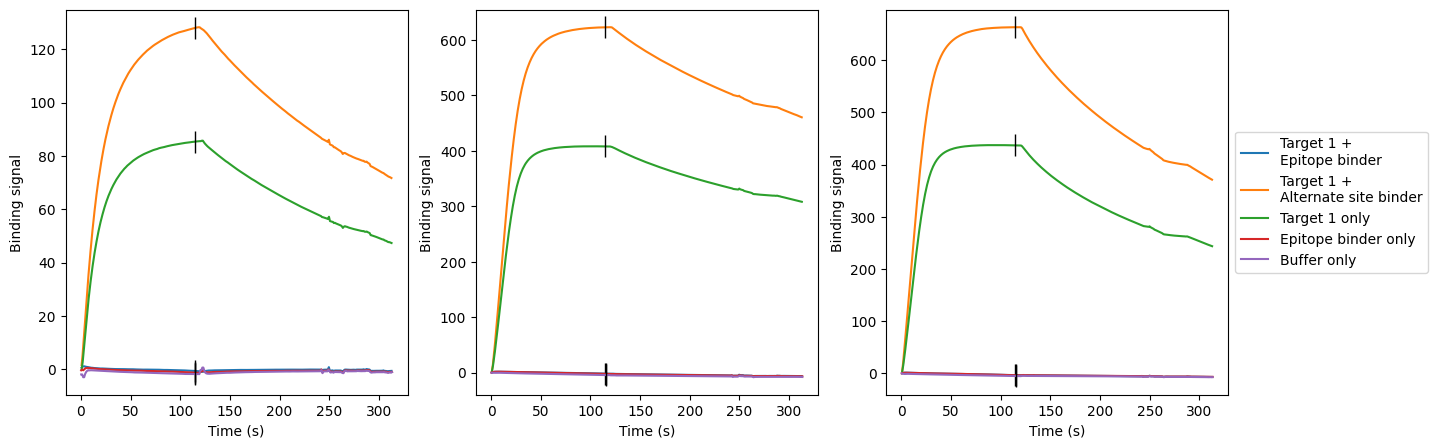
Figure 3: Example epitope characterization data. Surface plasmon resonance binding curves show that the designs bind the target (green), as well as the target pre-complexed with a known binder targeting an alternate epitope (orange). We see no binding for the target pre-complexed with a known binder against the epitope of interest (blue).
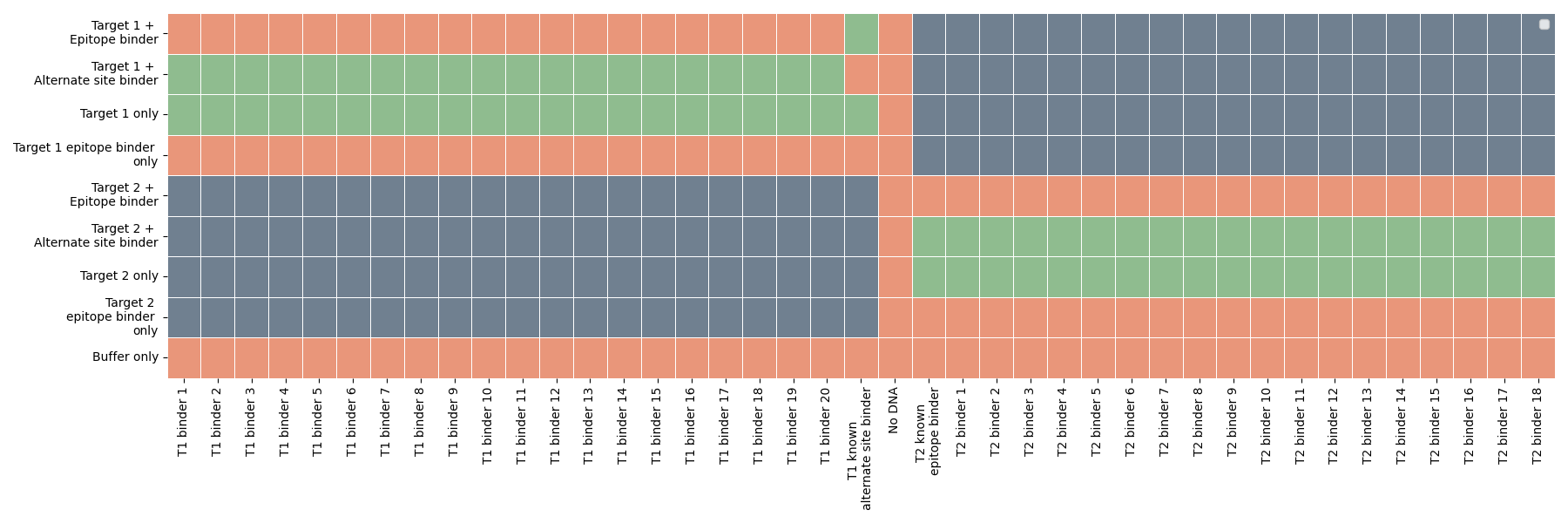
Figure 4: Epitope characterization data across a range of binders. Orange: no binding, Green: binding, Gray: not tested. All DSG-1 designs bind the target alone and the target pre-complexed with a second binder against an alternate epitope (green), and do not bind the target pre-complexed with a known binder against the epitope of interest (orange).
Summary
DSG-1 can design nanobody binders that bind specific target epitopes with a 3% hit rate. DSG-1 produces several strong binders (<=10nM binding) that are specific and target the correct epitope.